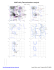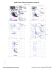MolProbity Ramachandran analysis
Transcrição
MolProbity Ramachandran analysis 2W49, model 1 General case 180 Isoleucine and valine 180 0 126 GLU 3 6 9 Z 2 141 LYS 5 8 2Z 8 112 ARG ARG 5 112 9 106 0 3 6 E P N JD L M S K IF G R H O Q LYS 233 233 233 233SER SER SER SER Y 1 246 4 7 246 LYS L E M S K IF G R P N JD H O Q 246 246 246 246GLN GLN GLN GLN Psi 2Z 5 8 0 Z 2 102 LEU 5 8 Z 2 136 LEU 5 8 Z 852 132 VAL Psi 9630 160 VAL 44 GLU GLU Y 1 188 4 7 188 ARG 9 62 GLU 0 3 6 Y 1 247 4 7 247 HIS 2 98 LYS 5 Z 8 V A 152 ASP 9630 107 ASN 0 L E M S K IF G R P N JD H O QZ 253 253 253 GLU GLU GLU 2253 5 8 7 GLU ARG L E M S K IF G R P N JD H O Q 2222GLU GLU GLU GLU L E S F D H O M K IG R P N JQ 274 274 274 274 274ILE ILE ILE ILE ILE V A9 0 151 3 6 30 ALA ALA Z 2Z 5 8 GLN 2 142 5 8 57 MET -180 -180 -180 0 Phi 180 Pre-proline 180 -180 Psi 0 0 Phi 180 Glycine 180 Psi 0 30 34 GLY 6 9 GLY Z 852 55 GLY L E M S K IG R P N JD H O Q F 242 242 242 242 242LEU LEU LEU LEU LEU -180 -180 -180 0 Phi 180 Trans proline 180 -180 0 Phi 180 Phi 180 Cis proline 180 Y 1 191 4 7 191 PRO Z 2 50 PRO 5 8 Psi Psi 0 0 Z 2 54 PRO 5 8 -180 -180 -180 89.2% (7764/8704) of all residues were in favored (98%) regions. F 246 GLN (-18.6, 73.4) 97.7% (8508/8704) of all residues were in allowed (>99.8%) regions. F 253 GLU (-40.5, -26.2) F 274 ILE (-23.1, -54.4) G 2 GLU (-25.3, -50.3) There were 196 outliers (phi, psi): 0 30 ALA (-58.1, -82.3) 0 34 GLY (162.0, 0.1) 0 62 GLU (-65.7, 15.5) 0 106 LYS (24.4, 93.2) 0 107 ASN (153.9, -12.3) 0 126 GLU (-28.7, 152.5) 0 160 VAL (-28.3, 106.6) 1 188 ARG (-62.8, 21.5) H 246 GLN (-18.8, 73.5) H 253 GLU (-40.7, -26.2) H 274 ILE (-23.0, -54.6) I 2 GLU (-25.5, -50.1) I 233 SER (47.9, 90.0) I 242 LEU (-65.1, -171.2) 54 PRO (-59.3, -160.9) 55 GLY (-59.4, -111.4) 57 MET (-67.7, -105.2) J 253 GLU (-40.7, -26.2) J 274 ILE (-22.9, -54.6) K 2 GLU (-25.5, -50.1) K 233 SER (47.9, 90.0) 3 34 GLY (161.9, 0.2) 3 62 GLU (-65.6, 15.3) 3 106 LYS (24.3, 93.1) K 242 LEU (-65.1, -171.2) K 246 GLN (-18.7, 73.5) K 253 GLU (-40.7, -26.2) 3 107 ASN (154.0, -12.2) 3 126 GLU (-28.7, 152.4) 3 160 VAL (-28.4, 106.5) K 274 ILE (-22.9, -54.6) L 2 GLU (-25.5, -50.3) L 233 SER (47.9, 89.9) 4 188 ARG (-63.0, 21.6) 4 191 PRO (-34.9, 157.7) 4 246 LYS (26.8, 78.7) 4 247 HIS (-60.4, 5.3) L 242 LEU (-65.1, -171.3) L 246 GLN (-18.7, 73.5) L 253 GLU (-40.5, -26.3) L 274 ILE (-23.1, -54.5) 5 5 5 M 2 GLU (-25.4, -50.3) M 233 SER (47.7, 90.0) M 242 LEU (-65.1, -171.2) 5 54 PRO (-59.3, -160.9) 5 55 GLY (-59.4, -111.5) 5 57 MET (-67.8, -105.1) 5 98 LYS (-49.3, -5.0) M 246 GLN (-18.7, 73.5) M 253 GLU (-40.6, -26.2) M 274 ILE (-23.0, -54.5) N 2 GLU (-25.4, -50.2) 5 102 LEU (-169.3, -11.8) 5 112 ARG (44.1, 99.0) 5 132 VAL (-156.4, 98.9) N 233 SER (47.8, 90.1) N 242 LEU (-65.2, -171.2) N 246 GLN (-18.8, 73.5) O 233 SER (47.8, 90.0) O 242 LEU (-65.1, -171.3) O 246 GLN (-18.8, 73.6) O 253 GLU (-40.6, -26.2) 6 107 ASN (154.0, -12.4) 6 126 GLU (-28.6, 152.5) 6 160 VAL (-28.2, 106.5) O 274 ILE (-23.0, -54.5) P 2 GLU (-25.6, -50.2) P 233 SER (47.7, 90.0) 7 188 ARG (-62.8, 21.4) 7 191 PRO (-34.8, 157.7) 7 246 LYS (26.7, 78.8) 7 247 HIS (-60.4, 5.3) P 242 LEU (-65.1, -171.2) P 246 GLN (-18.6, 73.4) P 253 GLU (-40.7, -26.1) P 274 ILE (-23.0, -54.6) 8 8 8 Q 2 GLU (-25.4, -50.3) Q 233 SER (47.9, 89.9) Q 242 LEU (-65.1, -171.3) 8 102 LEU (-169.3, -11.8) 8 112 ARG (44.1, 99.0) 8 132 VAL (-156.4, 98.9) 8 136 LEU (-172.6, -36.5) 8 141 LYS (-22.5, 126.1) 8 142 GLN (-68.3, -102.8) 9 30 ALA (-58.1, -82.3) 9 34 GLY (162.0, 0.2) 9 62 GLU (-65.6, 15.4) 9 106 LYS (24.4, 93.1) 0 N 253 GLU (-40.6, -26.2) N 274 ILE (-22.9, -54.6) O 2 GLU (-25.4, -50.3) 6 30 ALA (-58.2, -82.3) 6 34 GLY (161.9, 0.2) 6 62 GLU (-65.5, 15.3) 6 106 LYS (24.2, 93.1) 4 GLU (29.9, 43.2) 7 ARG (-36.0, -30.8) 50 PRO (-36.5, 99.7) -180 I 246 GLN (-18.7, 73.5) I 253 GLU (-40.7, -26.1) I 274 ILE (-23.0, -54.6) J 2 GLU (-25.6, -50.2) J 233 SER (47.7, 90.0) J 242 LEU (-65.1, -171.3) J 246 GLN (-18.8, 73.5) 2 136 LEU (-172.7, -36.4) 2 141 LYS (-22.4, 126.2) 2 142 GLN (-68.5, -102.8) 3 30 ALA (-58.0, -82.3) 8 54 PRO (-59.3, -160.9) 8 55 GLY (-59.3, -111.5) 8 57 MET (-67.7, -105.1) 8 98 LYS (-49.4, -5.0) 180 G 274 ILE (-22.9, -54.7) H 2 GLU (-25.5, -50.2) H 233 SER (47.9, 89.9) H 242 LEU (-65.2, -171.3) 4 GLU (29.9, 43.0) 7 ARG (-36.0, -30.8) 50 PRO (-36.7, 99.8) 2 2 2 2 98 LYS (-49.4, -5.0) 2 102 LEU (-169.4, -11.8) 2 112 ARG (43.9, 99.2) 2 132 VAL (-156.3, 99.0) 4 GLU (29.9, 43.2) 7 ARG (-35.9, -30.9) 50 PRO (-36.6, 99.7) Phi G 233 SER (47.8, 89.9) G 242 LEU (-65.2, -171.3) G 246 GLN (-18.8, 73.6) G 253 GLU (-40.7, -26.1) 1 191 PRO (-34.8, 157.7) 1 246 LYS (26.9, 78.7) 1 247 HIS (-60.3, 5.2) 2 2 2 5 136 LEU (-172.7, -36.5) 5 141 LYS (-22.5, 126.2) 5 142 GLN (-68.4, -102.8) 0 Q 246 GLN (-18.9, 73.6) Q 253 GLU (-40.6, -26.3) Q 274 ILE (-22.8, -54.8) R 2 GLU (-25.5, -50.1) R 233 SER (47.8, 90.0) R 242 LEU (-65.2, -171.2) R 246 GLN (-18.7, 73.5) R 253 GLU (-40.7, -26.2) R 274 ILE (-23.0, -54.6) S 2 GLU (-25.5, -50.3) S 233 SER (47.9, 89.9) S 242 LEU (-65.1, -171.3) S 246 GLN (-18.7, 73.6) S 253 GLU (-40.4, -26.3) 9 107 ASN (153.9, -12.4) 9 126 GLU (-28.6, 152.5) 9 160 VAL (-28.3, 106.5) S 274 ILE (-23.1, -54.5) V 151 ALA (-65.1, -81.8) V 152 ASP (-40.5, -10.9) A 151 ALA (-65.1, -81.7) A 152 ASP (-40.7, -10.8) D 2 GLU (-25.6, -50.2) D 233 SER (47.8, 90.0) Y 188 ARG (-62.7, 21.4) Y 191 PRO (-34.8, 157.7) Y 246 LYS (26.8, 78.7) Y 247 HIS (-60.4, 5.3) D 242 LEU (-65.3, -171.2) D 246 GLN (-18.6, 73.4) D 253 GLU (-40.6, -26.3) Z 4 GLU (30.0, 43.1) Z 7 ARG (-36.0, -30.8) Z 50 PRO (-36.6, 99.8) D 274 ILE (-23.1, -54.6) E 2 GLU (-25.4, -50.2) E 233 SER (47.6, 90.0) E 242 LEU (-65.2, -171.3) Z 54 PRO (-59.3, -160.9) Z 55 GLY (-59.4, -111.4) Z 57 MET (-67.6, -105.2) Z 98 LYS (-49.4, -5.0) E 246 GLN (-18.7, 73.5) E 253 GLU (-40.6, -26.2) E 274 ILE (-23.0, -54.6) Z 102 LEU (-169.4, -11.8) Z 112 ARG (44.0, 99.0) Z 132 VAL (-156.2, 99.0) F 2 GLU (-25.6, -50.1) F 233 SER (47.8, 90.0) F 242 LEU (-65.0, -171.3) Z 136 LEU (-172.7, -36.4) Z 141 LYS (-22.4, 126.3) Z 142 GLN (-68.5, -102.7) http://kinemage.biochem.duke.edu Lovell, Davis, et al. Proteins 50:437 (2003)
Documentos relacionados
MolProbity Ramachandran analysis
 97.8% (2392/2447) of all residues were in favored (98%) regions.
100.0% (2447/2447) of all residues were in allowed (>99.8%) regions.
There were no outliers.
97.8% (2392/2447) of all residues were in favored (98%) regions.
100.0% (2447/2447) of all residues were in allowed (>99.8%) regions.
There were no outliers.
MolProbity Ramachandran analysis
 3 ALA (87.5, -98.2)
4 ALA (171.2, 16.2)
6 GLY (-174.8, -58.2)
7 ASP (9.6, -102.1)
8 HIS (49.8, -18.1)
39 SER (-61.4, -124.3)
40 GLY (65.7, 129.7)
49 PRO (-59.4, 53.6)
8 ILE (78.9, 10.3)
10 ASN (88....
3 ALA (87.5, -98.2)
4 ALA (171.2, 16.2)
6 GLY (-174.8, -58.2)
7 ASP (9.6, -102.1)
8 HIS (49.8, -18.1)
39 SER (-61.4, -124.3)
40 GLY (65.7, 129.7)
49 PRO (-59.4, 53.6)
8 ILE (78.9, 10.3)
10 ASN (88....
MolProbity Ramachandran analysis
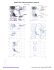 AZ 44 TRP (121.1, 138.3)
BB 24 LYS (-65.6, -165.2)
BB 27 VAL (-154.0, 34.4)
BB 41 LYS (-63.2, -167.9)
BB 52 ILE (-142.9, -107.8)
BB 53 PRO (-92.2, -148.8)
BB 95 TYR (-52.4, 91.0)
BB 113 GLU (131.9,...
AZ 44 TRP (121.1, 138.3)
BB 24 LYS (-65.6, -165.2)
BB 27 VAL (-154.0, 34.4)
BB 41 LYS (-63.2, -167.9)
BB 52 ILE (-142.9, -107.8)
BB 53 PRO (-92.2, -148.8)
BB 95 TYR (-52.4, 91.0)
BB 113 GLU (131.9,...
Ramachandran Plots
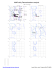 B 6 ALA (53.1, -88.5)
B 12 GLU (-66.3, -157.7)
B 13 ARG (38.0, -152.0)
B 19 ILE (-53.9, -79.5)
B 20 LYS (-28.0, -42.9)
B 37 LYS (-47.4, 90.3)
B 61 LEU (-59.1, -78.1)
B 71 GLN (-58.2, -86.0)
B 72 AS...
B 6 ALA (53.1, -88.5)
B 12 GLU (-66.3, -157.7)
B 13 ARG (38.0, -152.0)
B 19 ILE (-53.9, -79.5)
B 20 LYS (-28.0, -42.9)
B 37 LYS (-47.4, 90.3)
B 61 LEU (-59.1, -78.1)
B 71 GLN (-58.2, -86.0)
B 72 AS...
Ramachandran Plots
 75 THR (57.9, 107.4)
76 GLU (-176.0, -40.0)
78 ILE (-159.0, -42.6)
79 SER (60.6, 111.1)
84 GLN (59.3, 99.0)
85 VAL (59.3, 88.4)
89 LYS (-175.7, -40.1)
95 PRO (-53.5, -172.8)
96 LYS (-177.3, -39.6)
...
75 THR (57.9, 107.4)
76 GLU (-176.0, -40.0)
78 ILE (-159.0, -42.6)
79 SER (60.6, 111.1)
84 GLN (59.3, 99.0)
85 VAL (59.3, 88.4)
89 LYS (-175.7, -40.1)
95 PRO (-53.5, -172.8)
96 LYS (-177.3, -39.6)
...
MolProbity Ramachandran analysis
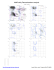 B 426 LYS (-61.4, -84.8)
B 427 ASP (-7.7, -71.3)
B 429 PHE (-54.8, 3.1)
B 509 ALA (82.5, 77.3)
B 510 LYS (-98.6, 59.8)
B 574 SER (-179.4, 52.3)
B 644 GLU (-173.6, -63.8)
B 645 SER (102.2, -104.5)
B...
B 426 LYS (-61.4, -84.8)
B 427 ASP (-7.7, -71.3)
B 429 PHE (-54.8, 3.1)
B 509 ALA (82.5, 77.3)
B 510 LYS (-98.6, 59.8)
B 574 SER (-179.4, 52.3)
B 644 GLU (-173.6, -63.8)
B 645 SER (102.2, -104.5)
B...
MolProbity Ramachandran analysis
 M 154 LEU (-50.8, -112.4)
M 162 ASP (17.6, -124.2)
M 164 ASP (44.7, 173.3)
M 181 ASP (164.8, 125.8)
M 183 ASP (42.2, 157.5)
N 20 SER (-47.3, -76.1)
M 154 LEU (-50.8, -112.4)
M 162 ASP (17.6, -124.2)
M 164 ASP (44.7, 173.3)
M 181 ASP (164.8, 125.8)
M 183 ASP (42.2, 157.5)
N 20 SER (-47.3, -76.1)
 BA1491
SER
AA1596
2112ASP
ALA
B 1302 ALA
BA1491
SER
AA1596
2112ASP
ALA
B 1302 ALA
 D
95 SER
B 352 ASN
A 352 ASN
D
95 SER
B 352 ASN
A 352 ASN
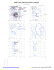 Pre-proline
Pre-proline
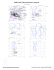 Phi
Phi
 Cis proline
Cis proline
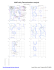
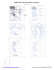 Pre-proline
Pre-proline
 D 488 GLY (-39.9, 153.4)
D 493 THR (-61.0, -173.9)
D 488 GLY (-39.9, 153.4)
D 493 THR (-61.0, -173.9)